|
9/21/2023 0 Comments Gene sequence definition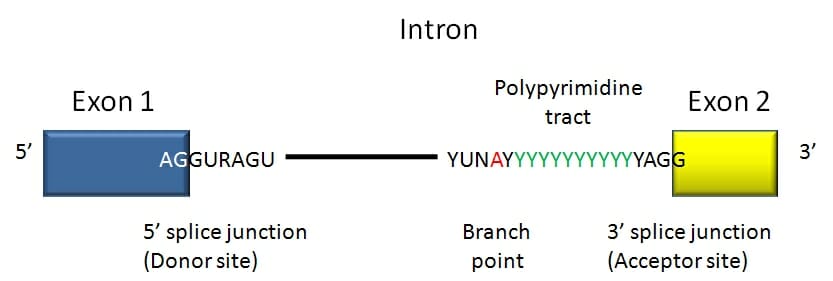
The output of Markov models in the context of annotation includes the probabilities of every kind of genomic element in every single part of the genome, and an accurate Markov model will assign high probabilities to correct annotations and low probabilities to the incorrect ones. To ensure a Markov model detects a genomic signal, it must first be trained on a series of known genomic signals. 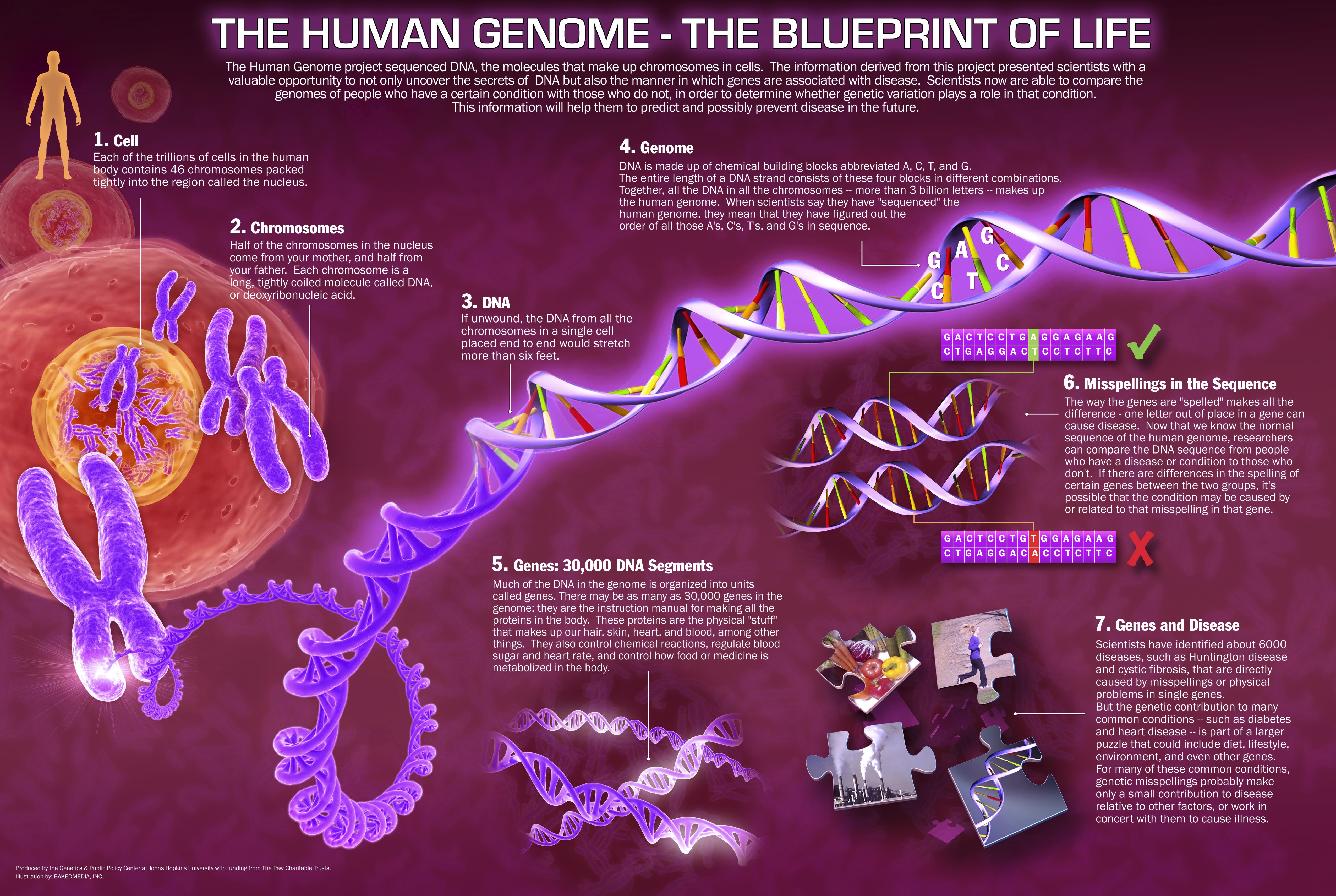
Markov models are the driving force behind many algorithms used within annotators of this generation these models can be thought of as directed graphs where nodes represent different genomic signals (such as transcription and translation start sites) connected by arrows representing the scanning of the sequence. Just like in the previous generation, they performed annotation through ab initio methods, but now applied on a genome-wide scale. The advent of complete genomes in the 1990s (the first one being the genome of Haemophilus influenzae sequenced in 1995) introduced a second generation of annotators. This was also known to be the case for synonymous codons, which are often present in proteins expressed at a lower level. In fact, codon usage was the main strategy used by several early protein coding sequence (CDS) prediction methods, based on the assumption that the most translated regions in a genome contain codons with the most abundant corresponding tRNAs (the molecules responsible for carrying amino acids to the ribosome during protein synthesis) allowing a more efficient translation. It performed several tasks related to annotation, such as base and codon counts. The first software used to analyze sequencing reads is the Staden Package, created by Rodger Staden in 1977. They appeared as a necessity to handle the enormous amount of data produced by the Maxam-Gilbert and Sanger DNA sequencing techniques developed in the late 1970s. The first generation of genome annotators used local ab initio methods, which are based solely on the information that can be extracted from the DNA sequence on a local scale, that is, one open reading frame (ORF) at a time. This is not the only way in which it has been categorized, as several alternatives, such as dimension-based and level-based classifications, have also been proposed. ĭNA annotation is classified into two categories: structural annotation, which identifies and demarcates elements in a genome, and functional annotation, which assigns functions to these elements. However, the conclusions drawn from the obtained results require manual expert analysis. Furthermore, due to the size and complexity of sequenced genomes, DNA annotation is not performed manually, but is instead automated by computational means. Although describing individual genes and their products or functions is sufficient to consider this description as an annotation, the depth of analysis reported in literature for different genomes vary widely, with some reports including additional information that goes beyond a simple annotation. Īnnotation is performed after a genome is sequenced and assembled, and is a necessary step in genome analysis before the sequence is deposited in a database and described in a published article. Among other things, it identifies the locations of genes and all the coding regions in a genome and determines what those genes do. In molecular biology and genetics, DNA annotation or genome annotation is the process of describing the structure and function of the components of a genome, by analyzing and interpreting them in order to extract their biological significance and understand the biological processes in which they participate. Represented with arrows, the transcription directions for the inner and outer genes are listed clockwise and anticlockwise, respectively. All individual genes are placed on the outermost circle according to their position in the genome, their transcription direction and their length they are color-coded based on the cellular function or component they are part of. The outer gray circle shows GC content in the every section of the genome. The number of genes, the genome length, and the GC content are placed in the middle black circle. The process of describing the structure and function of a genome A visualization of Porphyra umbilicalis chloroplast genome annotation ( GenBank accession: MF385003.1) made with Chloroplot.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
